Comparative Analysis
Vitessce can display the results of case-versus-control style comparative analyses of single-cell datasets (example).
There are not yet standard or widely-adopted approaches to store the results of comparative analyses. As noted by Hrovatin et al. Nature Methods 2024:
Moreover, there is a lack of consensus on where such custom marker gene and gene program lists could be deposited for easy querying across atlases. Thus, there is a need to define the type of analysis results that should ideally be provided by reference atlases, and to develop the necessary infrastructure to share these outputs.
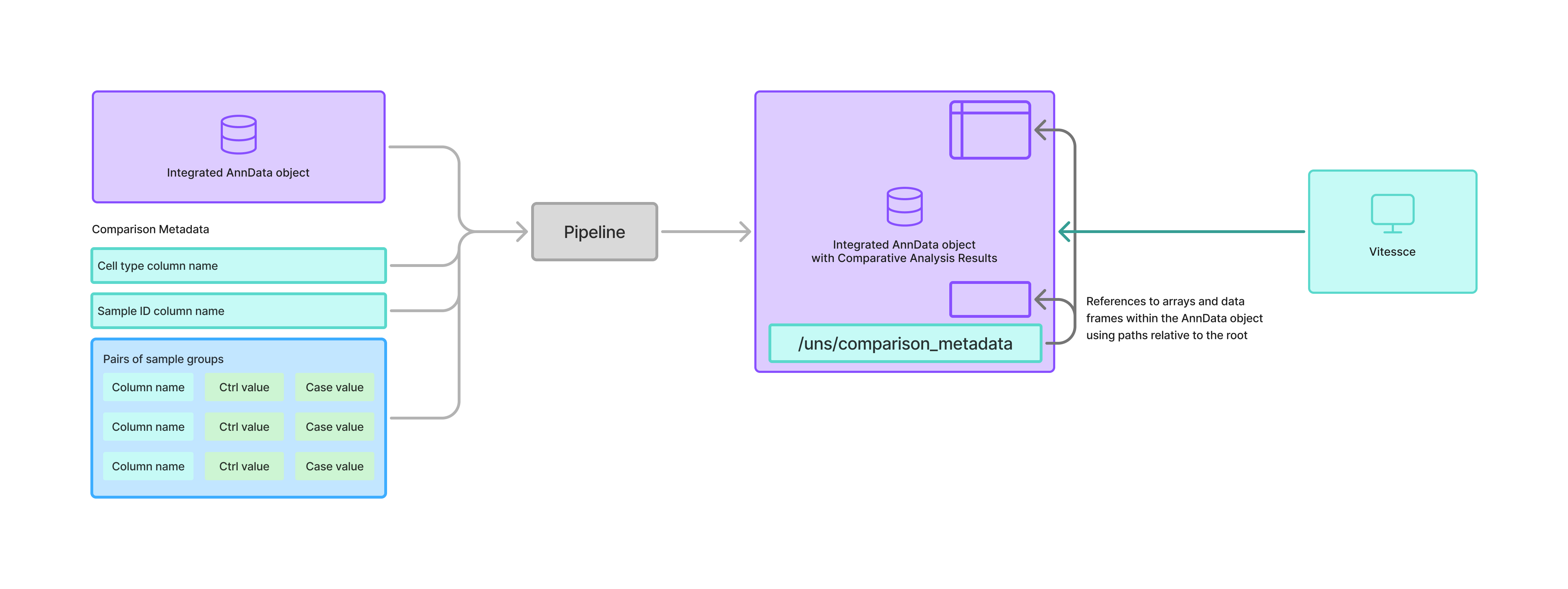
To address this problem, we have defined one such approach in which we store metadata about the comparative analyses that have been performed within the "unstructured" (uns) section of an AnnData object.
This metadata refers back to result-containing dataframes or arrays stored within the AnnData object.
The metadata format proposed here is in the early stages and subject to change.
We have defined JSON and Zod schemas to store this metadata.
We serialize the metadata to a string prior to storing in a subkey of uns (to avoid the otherwise recursive storage process for dictionary mappings) (e.g., z["uns/comparison_metadata"] = json.dumps(metadata_dict)).
A Python class can help with constructing such a metadata dict within data processing pipeline code.
At the top-level the dictionary contains the following properties:
schema_version(string): The version of the comparison metadata schema.cell_type_cols(string[]): Array of one or more columns inadata.obsstoring cell type annotations.sample_id_col(string): The column ofadata.obswhich maps cells to sample IDs.sample_group_pairs([string, [string, string]][]): Array of sample group pairs in the form of[colname, [ctrl_val, case_val]].comparisons({ [string]: { comparison: ComparisonPath, results: ComparisonResult[] } }): Dictionary mapping "comparison key"s to objects with the propertiescomparisonandresults. The value ofcomparisonshould be the "comparison path", a string or (nested) array of strings which represent the pre-slug-ified comparison key. See below for the expected format ofComparisonResultdictionaries. Note thatresultsstores an array, since the same comparison key/path may correspond to more than one comparison result (e.g., for the comparison keycompare_celltype.val_macrophage.__rest__, we might perform both a differential expression test and a gene set enrichment test, producing two separate comparison result dataframes).
Within each of the results entries, the dictionary contains the following properties:
path(string): The path to the dataframe, relative to the AnnData object root.coordination_values({ [string]: string }): Key-value pairs in the form of Vitessce coordination values (i.e.,coordinationType:coordinationValuemappings).analysis_type(string): The type of comparative analysis method which was applied to generate this result (e.g.,rank_genes_groupsorenrich).analysis_params(null|object): Parameters that are relevant to the analysis. This may take a different form depending on the analysis type.
We present an annotated (partial) example below:
Example
{
// The current schema version
"schema_version": "0.0.2",
// Cell type columns (of cell-level adata.obs dataframe)
"cell_type_cols": [
"cell_type",
"subclass_l1",
"subclass_l2"
],
// Sample ID column (of cell-level adata.obs dataframe)
"sample_id_col": "SampleID",
// Array of sample group (control, case) pairs
"sample_group_pairs": [
[
// Column name (of sample metadata dataframe)
"EnrollmentCategory",
// Tuple of (control, case) values in this column
[
"Reference", // Control value
"AKI" // Case value
]
],
// Additional [ column_name, [control_val, case_val] ] pairs
[ "EnrollmentCategory", ["AKI", "H-CKD"] ],
[ "EnrollmentCategory", ["DKD", "Reference"] ],
[ "AdjudicatedCategory", ["Diabetic Kidney Disease", "Reference"] ],
[ "AdjudicatedCategory", ["Acute Tubular Injury", "Reference"] ],
[ "AdjudicatedCategory", ["Acute Interstitial Nephritis", "Reference"] ],
[ "EnrollmentCategory", ["DKD", "H-CKD"] ],
[ "AdjudicatedCategory", ["Diabetic Kidney Disease", "Hypertensive Kidney Disease"] ],
[ "AdjudicatedCategory", ["Acute Interstitial Nephritis", "Acute Tubular Injury"] ],
[ "diseasetype", ["Reference", "AKI"] ],
[ "diseasetype", ["CKD", "AKI"] ],
[ "diseasetype", ["Reference", "CKD"] ]
],
// Donor ID column (optional, not yet used)
"donor_id_col": "donor_id",
// Metadata about comparisons that have been performed.
"comparisons": {
"__all__": { // Comparison key (an arbitrary string, but typically reflects the comparison path)
"comparison": "__all__", // Comparison path: string/array of strings/array of arrays of strings
"results": [ // Array of results for this comparison
{ // One comparative analysis result corresponding to this comparison path
"path": "uns/__all__.samples", // Path to a dataframe, relative to AnnData object root
"coordination_values": { // "Coordination values" which define the entities being compared.
"obsType": "sample" // <coordination type>: <coordination value> pairs
},
"analysis_type": "samples", // What type of analysis method produced these results?
"analysis_params": null // Place to store parameters used for this analysis (specific to the analysis_type).
}
]
},
"compare_cell_type.val_kidney_collecting_duct_principal_cell.__rest__": { // Another comparison key
"comparison": [["compare", "cell_type"], ["val", "kidney collecting duct principal cell"], "__rest__"],
"results": [
{
"path": "uns/compare_cell_type.val_kidney_collecting_duct_principal_cell.__rest__.rank_genes_groups",
"coordination_values": {
"obsType": "cell",
"featureType": "gene",
// For one cell type vs. the rest, we specify one [cell_type_column, cell_type_value] path.
"obsSetSelection": [
["cell_type", "kidney collecting duct principal cell"]
]
},
// Differential expression results from the ScanPy rank_genes_groups function.
"analysis_type": "rank_genes_groups",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type",
"reference": "rest",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"rank_genes_groups_df": {
"group": "kidney collecting duct principal cell"
}
}
},
{
"path": "uns/compare_cell_type.val_kidney_collecting_duct_principal_cell.__rest__.pertpy_hypergeometric",
"coordination_values": {
"obsType": "cell",
"featureType": "pathway",
"obsSetSelection": [
["cell_type","kidney collecting duct principal cell"]
]
},
// Gene set enrichment analysis results from the PertPy hypergeometric function.
"analysis_type": "pertpy_hypergeometric",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type",
"reference": "rest",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"pertpy_hypergeometric": {
"group": "kidney collecting duct principal cell",
"pvals_adj_thresh": 0.05,
"direction": "both",
"corr_method": "benjamini-hochberg"
}
}
},
{
"path": "uns/compare_cell_type.val_kidney_collecting_duct_principal_cell.__rest__.enrich",
"coordination_values": {
"obsType": "cell",
"featureType": "pathway",
"obsSetSelection": [
["cell_type", "kidney collecting duct principal cell"]
]
},
// Gene set enrichment analysis results from the ScanPy enrich function.
"analysis_type": "enrich",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type",
"reference": "rest",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"enrich": {
"group": "kidney collecting duct principal cell",
"log2fc_min": 2,
"pval_cutoff": 0.01
}
}
}
]
},
"compare_cell_type.val_kidney_interstitial_cell.__rest__": { // Another comparison key
"comparison": [["compare", "cell_type"], ["val", "kidney interstitial cell"], "__rest__"],
"results": [
{
"path": "uns/compare_cell_type.val_kidney_interstitial_cell.__rest__.rank_genes_groups",
"coordination_values": {
"obsType": "cell",
"featureType": "gene",
"obsSetSelection": [
["cell_type", "kidney interstitial cell"]
]
},
// Differential expression results from the ScanPy rank_genes_groups function.
"analysis_type": "rank_genes_groups",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type",
"reference": "rest",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"rank_genes_groups_df": {
"group": "kidney interstitial cell"
}
}
},
{
"path": "uns/compare_cell_type.val_kidney_interstitial_cell.__rest__.pertpy_hypergeometric",
"coordination_values": {
"obsType": "cell",
"featureType": "pathway",
"obsSetSelection": [
["cell_type", "kidney interstitial cell"]
]
},
// Gene set enrichment analysis results from the PertPy hypergeometric function.
"analysis_type": "pertpy_hypergeometric",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type",
"reference": "rest",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"pertpy_hypergeometric": {
"group": "kidney interstitial cell",
"pvals_adj_thresh": 0.05,
"direction": "both",
"corr_method": "benjamini-hochberg"
}
}
},
{
"path": "uns/compare_cell_type.val_kidney_interstitial_cell.__rest__.enrich",
"coordination_values": {
"obsType": "cell",
"featureType": "pathway",
"obsSetSelection": [
["cell_type", "kidney interstitial cell"]
]
},
// Gene set enrichment analysis results from the ScanPy enrich function.
"analysis_type": "enrich",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type",
"reference": "rest",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"enrich": {
"group": "kidney interstitial cell",
"log2fc_min": 2,
"pval_cutoff": 0.01
}
}
}
]
},
"compare_diseasetype.val_reference.val_ckd.compare_subclass_l2": { // Another comparison key
"comparison": [["compare", "diseasetype"], ["val", "Reference"], ["val", "CKD"], ["compare", "subclass_l2"]],
"results": [
{
"path": "uns/compare_diseasetype.val_reference.val_ckd.compare_subclass_l2.sccoda_df",
"coordination_values": {
"obsType": "cell",
"sampleSetSelection": [
["diseasetype", "CKD"]
],
// Which sample groups were included?
"sampleSetFilter": [
["diseasetype", "Reference"],
["diseasetype", "CKD"]
],
"obsSetSelection": [
// For scCODA, this should be one "path" to the cell type column used.
["subclass_l2"]
]
},
// ScCODA cell type composition results.
"analysis_type": "sccoda_df",
"analysis_params": {
// What cell type did scCODA automatically choose as the "reference" cell type.
"reference_cell_type": "MON",
"automatic_reference_absence_threshold": 0.05
}
}
]
},
"compare_adjudicatedcategory.val_diabetic_kidney_disease.val_reference.compare_cell_type": { // Another comparison key
"comparison": [["compare","AdjudicatedCategory"], ["val", "Diabetic Kidney Disease"], ["val", "Reference"], ["compare", "cell_type"]],
"results": [
{
"path": "uns/compare_adjudicatedcategory.val_diabetic_kidney_disease.val_reference.compare_cell_type.sccoda_df",
"coordination_values": {
"obsType": "cell",
"sampleSetSelection": [
["AdjudicatedCategory", "Reference"]
],
"sampleSetFilter": [
["AdjudicatedCategory", "Diabetic Kidney Disease"],
["AdjudicatedCategory", "Reference"]
],
"obsSetSelection": [
["cell_type"]
]
},
// ScCODA cell type composition results.
"analysis_type": "sccoda_df",
"analysis_params": {
"reference_cell_type": "conventional dendritic cell",
"automatic_reference_absence_threshold": 0.05
}
}
]
},
"filter_subclass_l2.val_dfib.compare_diseasetype.val_reference.val_ckd": { // Another comparison key
"comparison": [["filter", "subclass_l2"], ["val", "dFIB"], ["compare", "diseasetype"], ["val", "Reference"], ["val", "CKD"]],
"results": [
{
"path": "uns/filter_subclass_l2.val_dfib.compare_diseasetype.val_reference.val_ckd.rank_genes_groups",
"coordination_values": {
"obsType": "cell",
"featureType": "gene",
"obsSetFilter": [
// Which cell type was this analysis filtered to?
["subclass_l2", "dFIB"]
],
"sampleSetSelection": [
["diseasetype", "CKD"]
],
"sampleSetFilter": [
// Which sample groups was this analysis filtered to (and were being compared)?
["diseasetype", "Reference"],
["diseasetype", "CKD"]
]
},
// Differential expression results from the ScanPy rank_genes_groups function.
"analysis_type": "rank_genes_groups",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type_sample_group",
"reference": "dFIB_Reference",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"rank_genes_groups_df": {
"group": "dFIB_CKD"
}
}
},
{
"path": "uns/filter_subclass_l2.val_dfib.compare_diseasetype.val_reference.val_ckd.pertpy_hypergeometric",
"coordination_values": {
"obsType": "cell",
"featureType": "pathway",
"obsSetFilter": [
["subclass_l2", "dFIB"]
],
"sampleSetSelection": [
["diseasetype", "CKD"]
],
"sampleSetFilter": [
["diseasetype", "Reference"],
["diseasetype", "CKD"]
]
},
// Gene set enrichment analysis results from the PertPy hypergeometric function.
"analysis_type": "pertpy_hypergeometric",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type_sample_group",
"reference": "dFIB_Reference",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"pertpy_hypergeometric": {
"group": "dFIB_CKD",
"pvals_adj_thresh": 0.05,
"direction": "both",
"corr_method": "benjamini-hochberg"
}
}
},
{
"path": "uns/filter_subclass_l2.val_dfib.compare_diseasetype.val_reference.val_ckd.enrich",
"coordination_values": {
"obsType": "cell",
"featureType": "pathway",
"obsSetFilter": [
["subclass_l2", "dFIB"]
],
"sampleSetSelection": [
["diseasetype", "CKD"]
],
"sampleSetFilter": [
["diseasetype", "Reference"],
["diseasetype", "CKD"]
]
},
// Gene set enrichment analysis results from the ScanPy enrich function.
"analysis_type": "enrich",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type_sample_group",
"reference": "dFIB_Reference",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"enrich": {
"group": "dFIB_CKD",
"log2fc_min": 2,
"pval_cutoff": 0.01
}
}
}
]
},
"filter_subclass_l1.val_t.compare_diseasetype.val_ckd.val_aki": { // Another comparison key
"comparison": [["filter","subclass_l1"], ["val", "T"], ["compare", "diseasetype"], ["val", "CKD"], ["val", "AKI"]],
"results": [
{
"path": "uns/filter_subclass_l1.val_t.compare_diseasetype.val_ckd.val_aki.rank_genes_groups",
"coordination_values": {
"obsType": "cell",
"featureType": "gene",
"obsSetFilter": [
["subclass_l1", "T"]
],
"sampleSetSelection": [
["diseasetype", "AKI"]
],
"sampleSetFilter": [
["diseasetype", "CKD"],
["diseasetype", "AKI"]
]
},
// Differential expression results from the ScanPy rank_genes_groups function.
"analysis_type": "rank_genes_groups",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type_sample_group",
"reference": "T_CKD",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"rank_genes_groups_df": {
"group": "T_AKI"
}
}
},
{
"path": "uns/filter_subclass_l1.val_t.compare_diseasetype.val_ckd.val_aki.pertpy_hypergeometric",
"coordination_values": {
"obsType": "cell",
"featureType": "pathway",
"obsSetFilter": [
["subclass_l1", "T"]
],
"sampleSetSelection": [
["diseasetype", "AKI"]
],
"sampleSetFilter": [
["diseasetype", "CKD"],
["diseasetype", "AKI"]
]
},
// Gene set enrichment analysis results from the PertPy hypergeometric function.
"analysis_type": "pertpy_hypergeometric",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type_sample_group",
"reference": "T_CKD",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"pertpy_hypergeometric": {
"group": "T_AKI",
"pvals_adj_thresh": 0.05,
"direction": "both",
"corr_method": "benjamini-hochberg"
}
}
},
{
"path": "uns/filter_subclass_l1.val_t.compare_diseasetype.val_ckd.val_aki.enrich",
"coordination_values": {
"obsType": "cell",
"featureType": "pathway",
"obsSetFilter": [
["subclass_l1", "T"]
],
"sampleSetSelection": [
["diseasetype", "AKI"]
],
"sampleSetFilter": [
["diseasetype", "CKD"],
["diseasetype", "AKI"]
]
},
// Gene set enrichment analysis results from the ScanPy enrich function.
"analysis_type": "enrich",
"analysis_params": {
"rank_genes_groups": {
"groupby": "cell_type_sample_group",
"reference": "T_CKD",
"method": "wilcoxon",
"use_raw": false,
"layer": "logcounts",
"corr_method": "benjamini-hochberg"
},
"enrich": {
"group": "T_AKI",
"log2fc_min": 2,
"pval_cutoff": 0.01
}
}
}
]
}
}
}